General Information/Overview
|
Intracellular MGE or transposable elements (TE) include transposons (Tn) and insertion sequences (IS) but can embrace integrons (In) [3][4] and introns [5][6][7]. Originally Tn were distinguished from IS since they carry passenger (also called cargo) genes not involved in catalyzing or regulating TE movement. Most eukaryotic DNA transposons have relatives among the prokaryotic IS (see [8]) and it is not surprising that a variety of these elements carrying passenger genes are now also being identified [9][10]. Prokaryotes harbor a host of such elements as well as several types of structure possessing characteristics of both groups (e.g. Integrative Conjugative Elements, ICE, originally called conjugative transposons, as well as other types of non-conjunctive genomic islands) [11][12][13].
|
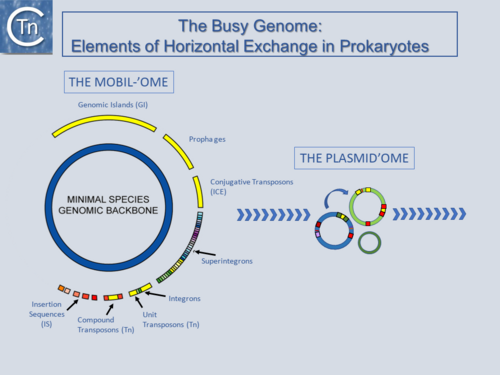 Fig.1.1.1. The Busy Genome: Elements of Horizontal Exchange. The genome backbone, which includes housekeeping genes, is shown as the inner circle (blue). The "mobilome" is shown in the outer circle. This includes a number of different types of MGE both intercellular (some genomic islands, prophages, and conjugative transposons) and intracellular (Insertion sequences, compound and unit transposons, integrons, and super integrons). An important class of intercellular MGE, the plasmids, act as transposon vectors and facilitate TE movement within the plasmidome. |
Bibliography
- ↑ </nowiki>
- ↑ </nowiki>
- ↑ </nowiki>
- ↑ </nowiki>
- ↑ </nowiki>
- ↑ </nowiki>
- ↑ </nowiki>
- ↑ Hickman AB, Chandler M, Dyda F . Integrating prokaryotes and eukaryotes: DNA transposases in light of structure. - Crit Rev Biochem Mol Biol: 2010 Feb, 45(1);50-69 [PubMed:20067338] [DOI] </nowiki>
- ↑ </nowiki>
- ↑ Bao W, Jurka J . Homologues of bacterial TnpB_IS605 are widespread in diverse eukaryotic transposable elements. - Mob DNA: 2013 Apr 1, 4(1);12 [PubMed:23548000] [DOI] </nowiki>
- ↑ </nowiki>
- ↑ Dobrindt U, Hochhut B, Hentschel U, Hacker J . Genomic islands in pathogenic and environmental microorganisms. - Nat Rev Microbiol: 2004 May, 2(5);414-24 [PubMed:15100694] [DOI] </nowiki>
- ↑ </nowiki>