Difference between revisions of "General Information/Influence of transposition mechanisms on genome impact"
Francislon (talk | contribs) m (Text replacement - "http://tncentral.ncc.unesp.br/cgi-bin/tn_report.pl?id=" to "https://tncentral.ncc.unesp.br/report/te/") |
|||
| Line 1: | Line 1: | ||
'''<big>T</big>'''he way in which strand cleavages and transfers occur during transposition also affects the outcome of the transposition events and therefore impinges on genome structure. For IS with DDE Tpases, [[Transposons families/Tn3 family|Tn''3'']] and [[IS Families/IS6 family|IS''6'' family]] members generate fusions or cointegrates between the donor and target replicons by a process of replicative transposition, presumably by '''T'''arget '''P'''rimed '''R'''eplicative '''T'''ransposition ('''TPRT''')<ref><pubmed>20067338</pubmed></ref><ref><pubmed>26104718</pubmed></ref><ref><pubmed>26104715</pubmed></ref> [[:Image:1.23.1.png|(Fig.17.1]] and [[:Image:1.23.2.png|Fig.17.2 top)]]. | '''<big>T</big>'''he way in which strand cleavages and transfers occur during transposition also affects the outcome of the transposition events and therefore impinges on genome structure. For IS with DDE Tpases, [[Transposons families/Tn3 family|Tn''3'']] and [[IS Families/IS6 family|IS''6'' family]] members generate fusions or cointegrates between the donor and target replicons by a process of replicative transposition, presumably by '''T'''arget '''P'''rimed '''R'''eplicative '''T'''ransposition ('''TPRT''')<ref><pubmed>20067338</pubmed></ref><ref><pubmed>26104718</pubmed></ref><ref><pubmed>26104715</pubmed></ref> [[:Image:1.23.1.png|(Fig.17.1]] and [[:Image:1.23.2.png|Fig.17.2 top)]]. | ||
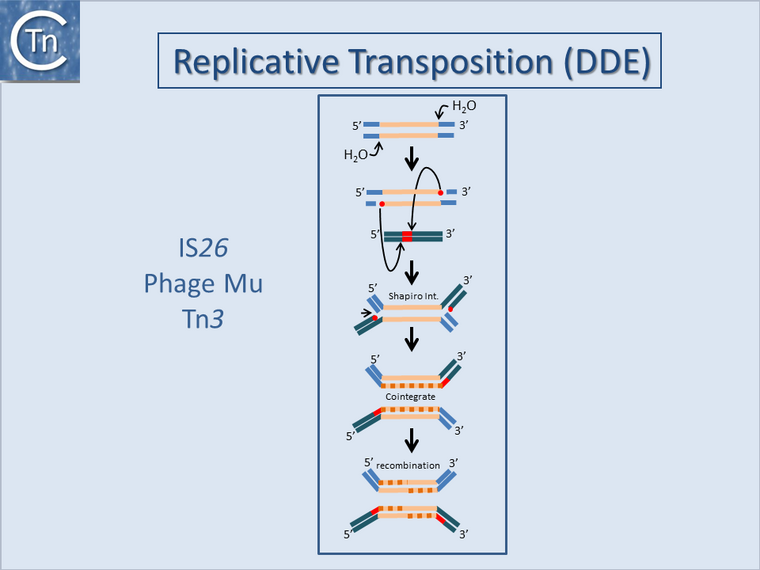
| − | However, in the event of intramolecular transposition, this type of mechanism is expected to give rise to inversions with a copy of the IS at each junction or inversions with a single IS copy remaining and a second copy segregating with a circularized deletion<ref><pubmed>287033</pubmed></ref> [[:Image:1.23.2.png|(Fig.17.2 botton]]; see <ref><pubmed>26060276</pubmed></ref>). Note that similar effects are also known to occur by [[wikipedia:Homologous_recombination|homologous recombination]] between two inverted or directly repeated IS copies in a replicon. Other known mechanisms such as cut-and-paste, or copy-and-paste ('''D'''onor '''P'''rimed '''R'''eplicatice '''T'''ransposition; '''DPRT''')<ref><pubmed>14682279</pubmed></ref> would not generate this type of genomic rearrangement but could contribute to genomic modifications in other ways such as “nearly precise excision”<ref><pubmed>455447</pubmed></ref> or by using alternative sequences which resemble their IR<ref><pubmed>6244582</pubmed></ref><ref><pubmed>8106332</pubmed></ref>.<br />[[Image:1.23.1.png|thumb|center|760x760px|'''Fig.17.1.''' Replicative Transposition ('''DDE''') of TE such as [ | + | However, in the event of intramolecular transposition, this type of mechanism is expected to give rise to inversions with a copy of the IS at each junction or inversions with a single IS copy remaining and a second copy segregating with a circularized deletion<ref><pubmed>287033</pubmed></ref> [[:Image:1.23.2.png|(Fig.17.2 botton]]; see <ref><pubmed>26060276</pubmed></ref>). Note that similar effects are also known to occur by [[wikipedia:Homologous_recombination|homologous recombination]] between two inverted or directly repeated IS copies in a replicon. Other known mechanisms such as cut-and-paste, or copy-and-paste ('''D'''onor '''P'''rimed '''R'''eplicatice '''T'''ransposition; '''DPRT''')<ref><pubmed>14682279</pubmed></ref> would not generate this type of genomic rearrangement but could contribute to genomic modifications in other ways such as “nearly precise excision”<ref><pubmed>455447</pubmed></ref> or by using alternative sequences which resemble their IR<ref><pubmed>6244582</pubmed></ref><ref><pubmed>8106332</pubmed></ref>.<br />[[Image:1.23.1.png|thumb|center|760x760px|'''Fig.17.1.''' Replicative Transposition ('''DDE''') of TE such as [https://tncentral.ncc.unesp.br/report/te/Tn3-V00613 Tn''3''] and [[wikipedia:Bacteriophage_Mu|bacteriophage Mu]]. The figure shows the transposition mechanism of replicative TE that uses a DDE Tpase. The transposon is represented as a yellow line. Flanking sequences in the donor molecule is blue. Flanking sequences in the target molecule are green. Red circles indicate 3′OH moieties generated by Tpase-catalyzed hydrolysis at the transposon end(s). Red boxes indicate target DNA flanks that are duplicated on insertion. '''Top to bottom:''' |
Tpase catalyzed cleavage at the 3′ transposon ends using H2O as the nucleophile. Liberated 3′OH attack the target in a staggered manner to create a branched molecule in which the transposon bridges both donor and target molecules. Replication proceeds, probably using the 3′OH liberated in the flanking target DNA, to generate a second transposon copy (newly replicated DNA is shown as a dotted orange line). If the donor and target DNA are circular molecules, this fuses the two, resulting in a cointegrate where the | Tpase catalyzed cleavage at the 3′ transposon ends using H2O as the nucleophile. Liberated 3′OH attack the target in a staggered manner to create a branched molecule in which the transposon bridges both donor and target molecules. Replication proceeds, probably using the 3′OH liberated in the flanking target DNA, to generate a second transposon copy (newly replicated DNA is shown as a dotted orange line). If the donor and target DNA are circular molecules, this fuses the two, resulting in a cointegrate where the | ||
Latest revision as of 05:06, 1 April 2024
The way in which strand cleavages and transfers occur during transposition also affects the outcome of the transposition events and therefore impinges on genome structure. For IS with DDE Tpases, Tn3 and IS6 family members generate fusions or cointegrates between the donor and target replicons by a process of replicative transposition, presumably by Target Primed Replicative Transposition (TPRT)[1][2][3] (Fig.17.1 and Fig.17.2 top).
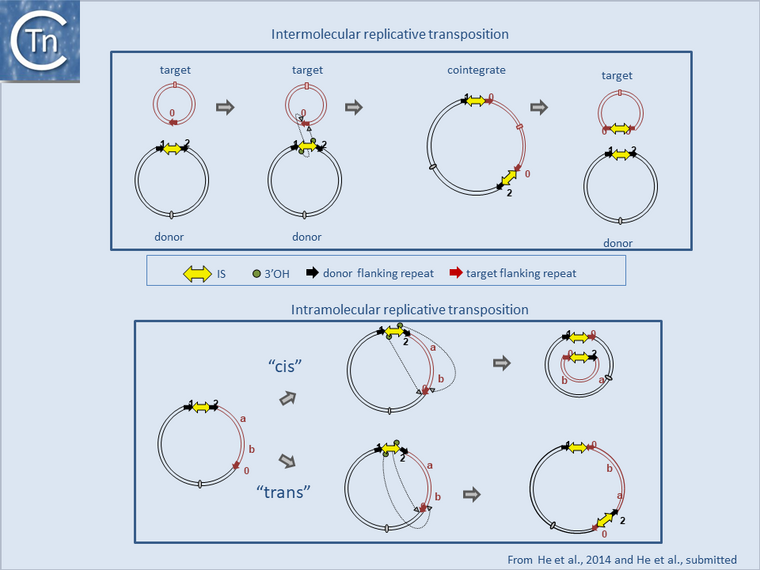
However, in the event of intramolecular transposition, this type of mechanism is expected to give rise to inversions with a copy of the IS at each junction or inversions with a single IS copy remaining and a second copy segregating with a circularized deletion[4] (Fig.17.2 botton; see [5]). Note that similar effects are also known to occur by homologous recombination between two inverted or directly repeated IS copies in a replicon. Other known mechanisms such as cut-and-paste, or copy-and-paste (Donor Primed Replicatice Transposition; DPRT)[6] would not generate this type of genomic rearrangement but could contribute to genomic modifications in other ways such as “nearly precise excision”[7] or by using alternative sequences which resemble their IR[8][9].
Bibliography
- ↑ Hickman AB, Chandler M, Dyda F . Integrating prokaryotes and eukaryotes: DNA transposases in light of structure. - Crit Rev Biochem Mol Biol: 2010 Feb, 45(1);50-69 [PubMed:20067338] [DOI]
- ↑
- ↑ Siguier P, Gourbeyre E, Varani A, Ton-Hoang B, Chandler M . Everyman's Guide to Bacterial Insertion Sequences. - Microbiol Spectr: 2015 Apr, 3(2);MDNA3-0030-2014 [PubMed:26104715] [DOI]
- ↑
- ↑ He S, Hickman AB, Varani AM, Siguier P, Chandler M, Dekker JP, Dyda F . Insertion Sequence IS26 Reorganizes Plasmids in Clinically Isolated Multidrug-Resistant Bacteria by Replicative Transposition. - mBio: 2015 Jun 9, 6(3);e00762 [PubMed:26060276] [DOI]
- ↑
- ↑ Ross DG, Swan J, Kleckner N . Nearly precise excision: a new type of DNA alteration associated with the translocatable element Tn10. - Cell: 1979 Apr, 16(4);733-8 [PubMed:455447] [DOI]
- ↑
- ↑ Polard P, Seroude L, Fayet O, Prère MF, Chandler M . One-ended insertion of IS911. - J Bacteriol: 1994 Feb, 176(4);1192-6 [PubMed:8106332] [DOI]