Difference between revisions of "General Information/Fuzzy Borders"
Jump to navigation
Jump to search
| (14 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
==Fuzzy Borders== | ==Fuzzy Borders== | ||
| − | '''<big>W</big>'''ith our increasing knowledge of the diversity of the mobilome, the distinction between IS and other TE is becoming increasing unclear. The major feature used to distinguish IS from transposons was that the former [[:Image:1.13.1.png|(Fig. | + | {| |
| + | |'''<big>W</big>'''ith our increasing knowledge of the diversity of the mobilome, the distinction between IS and other TE is becoming increasing unclear. The major feature used to distinguish IS from transposons was that the former [[:Image:1.13.1.png|(Fig.8.1)]] lack phenotypically detectable passenger genes (genes not involved in the transposition process) while the latter include one or more such genes (for antibiotic resistance, virulence and pathogenicity functions or genes permitting the use of unusual compounds). This is no longer the case [[:Image:1.13.1.png|(Fig.8.1)]]. | ||
| − | [[Image:1.13.1.png|thumb| | + | As shown in [[:Image:1.13.1.png|Fig.8.1]], examples have now been identified in which passenger genes are located within an IS, called tIS (ISs and relatives with passenger genes)<ref><pubmed>19286454</pubmed></ref> or in which TE with typical transposon structures are devoid of transposition proteins MITEs ('''M'''iniature '''I'''nverted repeat '''T'''ransposable '''E'''lements) and MICs ('''M'''obile '''I'''nsertion '''C'''assette))<ref><pubmed>15228527</pubmed></ref><ref><pubmed>10320586</pubmed></ref><ref><pubmed>11814653</pubmed></ref><ref><pubmed>12520302</pubmed></ref>. The (hypothetical) relationship between these different autonomous (with transposase) and non-autonomous IS derivatives is indicated in [[:Image:1.13.1.png|Fig.8.1]] <ref><pubmed>24499397</pubmed></ref><ref><pubmed>26104715</pubmed></ref>.<br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> |
| + | |||
| + | |[[Image:1.13.1.png|thumb|480x480px|'''Fig.8.1.''' Relationship between IS, tIS, and MITES. The (hypothetical) relationship between different IS derivatives (shown as pink boxes). Horizontal arrows indicate open reading frames encoding the Tpase (purple), passenger genes (red). The terminal inverted repeats are shown as dark blue triangles. Examples of IS families which include such derivatives are: [[IS Families/IS4 and related families#IS231|IS''4'' grp IS''231'']], [[IS Families/IS1595 family|IS''1595'']], and [[IS Families/IS66 family|IS''66'']].|alt=|none]] | ||
| + | |} | ||
==Bibliography== | ==Bibliography== | ||
| − | < | + | {{Reflist|32em}} |
| + | <br /> | ||
| + | <hr> | ||
| + | {{TnPedia}} | ||
Latest revision as of 12:36, 5 December 2022
Fuzzy Borders
| With our increasing knowledge of the diversity of the mobilome, the distinction between IS and other TE is becoming increasing unclear. The major feature used to distinguish IS from transposons was that the former (Fig.8.1) lack phenotypically detectable passenger genes (genes not involved in the transposition process) while the latter include one or more such genes (for antibiotic resistance, virulence and pathogenicity functions or genes permitting the use of unusual compounds). This is no longer the case (Fig.8.1).
As shown in Fig.8.1, examples have now been identified in which passenger genes are located within an IS, called tIS (ISs and relatives with passenger genes)[1] or in which TE with typical transposon structures are devoid of transposition proteins MITEs (Miniature Inverted repeat Transposable Elements) and MICs (Mobile Insertion Cassette))[2][3][4][5]. The (hypothetical) relationship between these different autonomous (with transposase) and non-autonomous IS derivatives is indicated in Fig.8.1 [6][7]. |
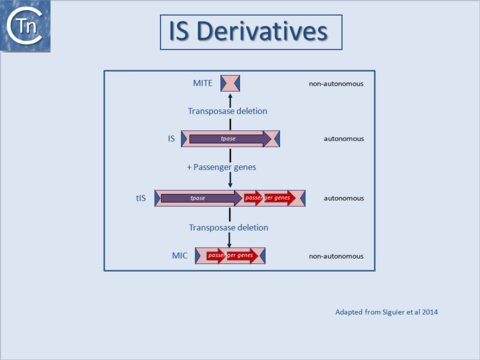 Fig.8.1. Relationship between IS, tIS, and MITES. The (hypothetical) relationship between different IS derivatives (shown as pink boxes). Horizontal arrows indicate open reading frames encoding the Tpase (purple), passenger genes (red). The terminal inverted repeats are shown as dark blue triangles. Examples of IS families which include such derivatives are: IS4 grp IS231, IS1595, and IS66. |
Bibliography
- ↑ Siguier P, Gagnevin L, Chandler M . The new IS1595 family, its relation to IS1 and the frontier between insertion sequences and transposons. - Res Microbiol: 2009 Apr, 160(3);232-41 [PubMed:19286454] [DOI]
- ↑ De Palmenaer D, Vermeiren C, Mahillon J . IS231-MIC231 elements from Bacillus cereus sensu lato are modular. - Mol Microbiol: 2004 Jul, 53(2);457-67 [PubMed:15228527] [DOI]
- ↑ Chen Y, Braathen P, Léonard C, Mahillon J . MIC231, a naturally occurring mobile insertion cassette from Bacillus cereus. - Mol Microbiol: 1999 May, 32(3);657-68 [PubMed:10320586] [DOI]
- ↑ Brügger K, Redder P, She Q, Confalonieri F, Zivanovic Y, Garrett RA . Mobile elements in archaeal genomes. - FEMS Microbiol Lett: 2002 Jan 10, 206(2);131-41 [PubMed:11814653] [DOI]
- ↑ Jiang N, Bao Z, Zhang X, Hirochika H, Eddy SR, McCouch SR, Wessler SR . An active DNA transposon family in rice. - Nature: 2003 Jan 9, 421(6919);163-7 [PubMed:12520302] [DOI]
- ↑ Siguier P, Gourbeyre E, Chandler M . Bacterial insertion sequences: their genomic impact and diversity. - FEMS Microbiol Rev: 2014 Sep, 38(5);865-91 [PubMed:24499397] [DOI]
- ↑ Siguier P, Gourbeyre E, Varani A, Ton-Hoang B, Chandler M . Everyman's Guide to Bacterial Insertion Sequences. - Microbiol Spectr: 2015 Apr, 3(2);MDNA3-0030-2014 [PubMed:26104715] [DOI]